|
The Metscape main window has three (3) tabs that provide users with the following options:ġ) Load a list of compounds (compound names or KEGG IDs) Ģ) Load a file containing normalized experimental metabolite data with metabolite KEGG IDs and corresponding values at given time points or under specific experimental conditions andģ) Display pathway-specific networks by choosing a pathway from the drop-down list. To start-up Metscape, users select Metscape from the plug-ins menu in Cytoscape. Users can easily apply Cytoscape layouts through the Cytoscape layout menu to the compound network generated by Metscape. 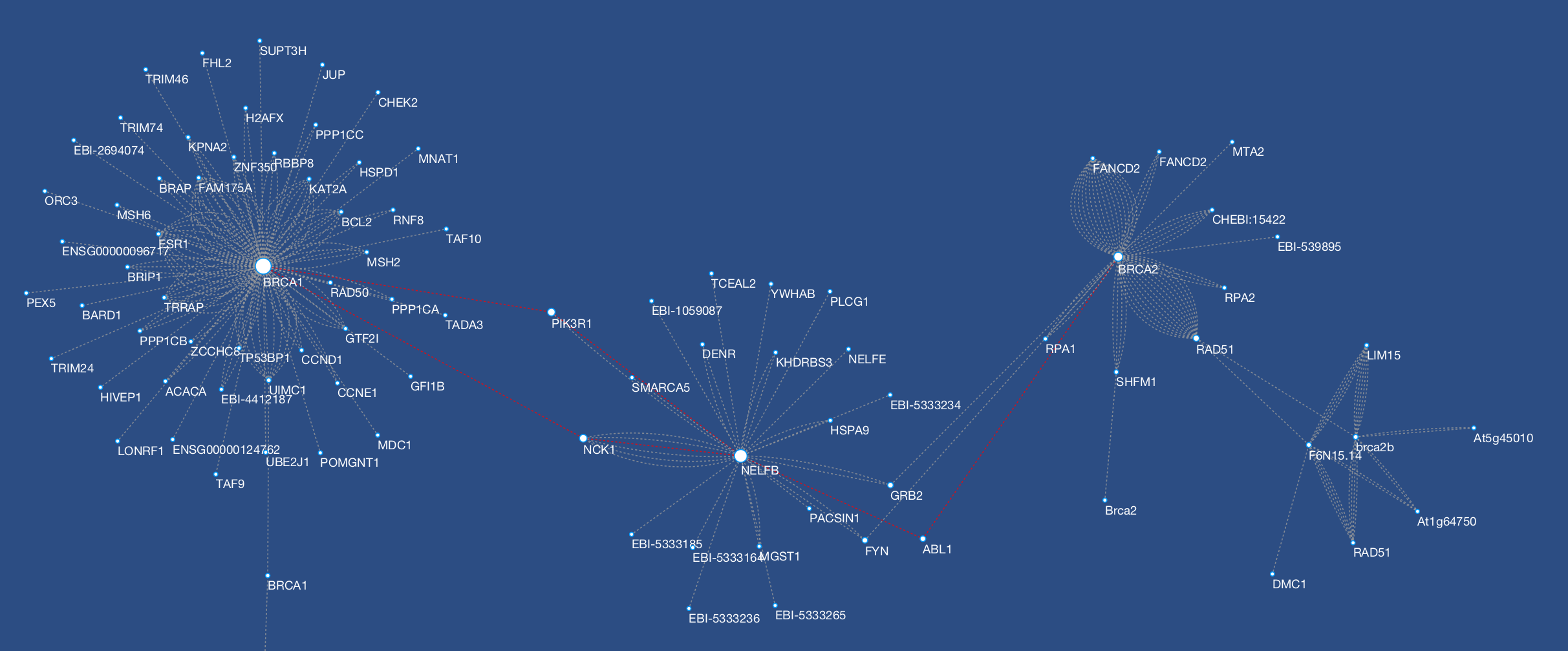
A number of different layout algorithms had been developed for Cytoscape that attempt to minimize the crossings between the edges, the distances between nodes and the bending of the edges. Typically, metabolic networks possess a high degree of connectivity. Visualization of metabolomic networks poses a number of challenges. MetScape uses an internal relational database stored at the National Center for Integrative Biomedical Informatics (NCIBI) that integrates data from the Kyoto Encyclopedia of Genes and Genomes (KEGG), the Edinburgh Human Metabolic Network (EHMN), and the NCIBI HUMDB Database. Gene expression and/or compound concentration data can be loaded from file(s), or the user can directly enter individual compounds/genes (using KEGG compound IDs, or Entrez Gene IDs) to build metabolic networks without loading a file. 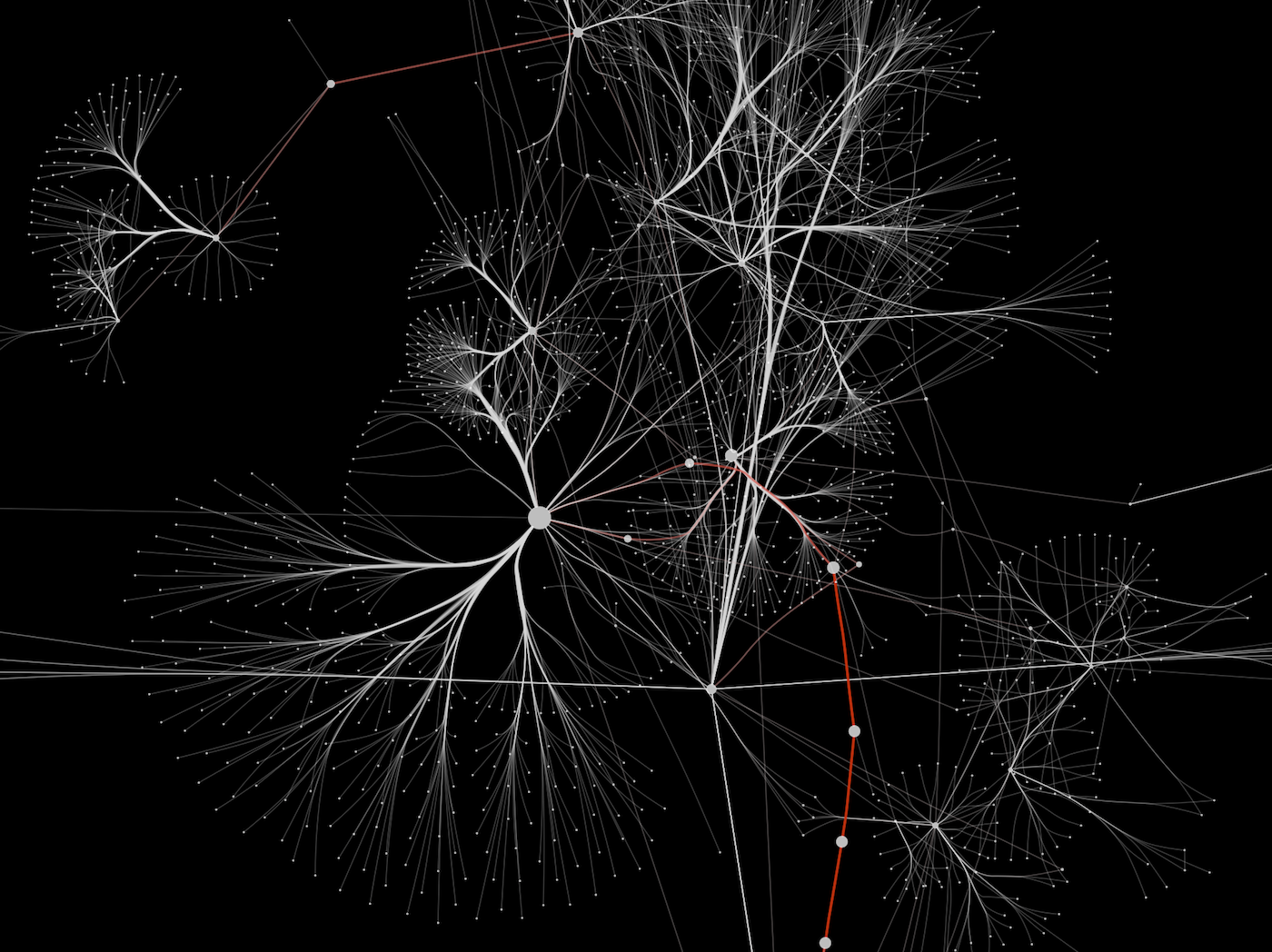
It allows users to build and analyze networks of genes and compounds, identify enriched pathways from expression profiling data, and visualize changes in metabolite data. Category Metabolomics/Metabonomics>Metabolic Profiling/Analysis Systems/Tools and Genomics>Gene Expression Analysis/Profiling/ToolsĪbstract MetScape Plugin for Cytoscape provides a bioinformatics framework for the visualization and interpretation of metabolomic and expression profiling data in the context of human metabolism.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
